Image Credit: Michael Freitag
Neurospora crassa is a filamentous fungus, i.e. growing as hyphae rather than forming unicellular yeast cells. Filamentous fungi are important in natural environments as degraders of plant and animal biomass, as pathogens of plants and animals, and as producers of natural products that are used as pharmaceuticals. The genus Neurospora holds a central position in the history of genetics, biochemistry, and molecular biology, as one of the earliest convenient model organisms that helped to decipher the gene to protein relationships of metabolism and gene regulation. Ongoing studies on light regulation, the circadian clock, chromatin and epigenetics, gene regulation, and metabolism continue to yield insights into general eukaryotic biology.
Neurospora crassa strain FGSC2225 Mauriceville-1cA, often called simply “Mauriceville” and abbreviated “MV” is a wild-collected strain from Texas, USA, and only distantly related to the reference strain FGSC2489 (aka “OR74A”). Mapping crosses between mutated strains derived from OR74A and MV were used to map mutations by restriction fragment length polymorphisms (1, 2) and more recently by Bulk Segregant Analyses followed by high-throughput sequencing (BSA-seq; ref. 3). Compared to the widely used reference and near-isogenic strains in the OR74A lineage, MV contains many large-scale changes in constitutively silent heterochromatin, and a divergent set of transposon relics (4).
References:
- Metzenberg RL, Stevens JN, Selker EU, Morzycka-Wroblewska E. Identification and chromosomal distribution of 5S rRNA genes in Neurospora crassa. Proceedings of the National Academy of Sciences of the United States of America. 82: 2067-71. PMID 3157192 DOI: 10.1073/PNAS.82.7.2067
- Nelson, M. A., and D.D. Perkins Restriction polymorphism maps of Neurospora crassa: 2000 update," Fungal Genetics Reports 2000 Vol. 47, Article 4. DOI: 10.4148/1941-4765.1200
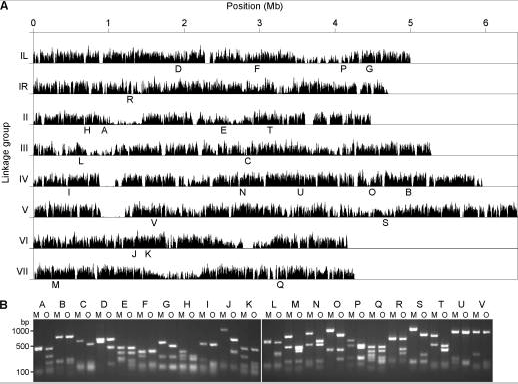
- Pomraning KR, Smith KM, Freitag M. Bulk segregant analysis followed by high-throughput sequencing reveals the Neurospora cell cycle gene, ndc-1, to be allelic with the gene for ornithine decarboxylase, spe-1. Eukaryotic Cell 2011 Jun;10(6):724-33. DOI: 10.1128/EC.00016-11. Epub 2011 Apr 22.
- Selker EU, Tountas NA, Cross SH, Margolin BS, Murphy JG, Bird AP, Freitag M. The methylated component of the Neurospora crassa genome. Nature. 422: 893-7. PMID 12712205 DOI: 10.1038/Nature01564
