| The Transcript Annotation area shows existing annotation information for the gene model and allows you to edit or add information. The name of the current transcript appears just below the Transcript Annotation heading. Beside it is a Hide or Show button.
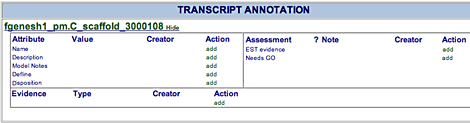
To view the Protein Page for this transcript, click the name of the transcript.
To hide all the transcript annotation data, click the Hide button.
To show all the transcript annotation data, click the Show button.
To add a transcript annotation (as an attribute, assessment, or evidence) click the green add button after it. This opens a small window with input fields for entering your annotation, usually just a text field plus Save Item and Cancel options. Enter your annotation data and then click Save Item. The data you entered appears in the Value, "?", Note, or Type column and your login ID appears in the Creator column. You may have to reload the web page to see the new annotation.
To edit an existing annotation, click edit. A window showing the current annotation opens. Make your changes to the input fields and click Save Item. You may have to reload the web page to see the change to the annotation.
To remove an annotation, click edit. In the window that opens, click Delete Item. You may have to reload the web page to see that the annotation has been deleted.
Annotatable items available in the Transcript Annotation box include
- Name The name you want to assign, usually based on the name of a gene in the same or another organism with very strong evidence of homology (e.g., via good BLAST alignment) or following a naming convention agreed upon by the annotators for this genome.
- Description A description of the function of the gene product.
- Model Notes Free text and/or notes from a controlled comment set (selected from a drop-down menu). To use both, add your free text after selecting a controlled comment from the drop-down menu.
- Defline Text in NCBI format (without the >) that can be used as the defline for the gene sequence, usually taken from the defline for a similar gene.
- Disposition An indication of the degree of confidence the annotator has in the gene model and whether it should be included in the catalog.
The Gene Catalog
The Gene Catalog is a working track for annotation purposes. Gene models from the Filtered Gene Models track (which includes gene models selected by hand as the best representative model available for each gene) are automatically added to the catalog initially. From the Disposition menu, annotators can then select
- Catalog to suggest a gene not already in the catalog be added to it,
- Tentative to suggest a gene model be added to the catalog on a tentative basis
- Delete to suggest the gene model be deleted from the catalog.
When editing a disposition annotation, an annotator can also directly delete the gene from the catalog by clicking the Delete Item button. (This is best done only when other annotators have suggested deletion.)
- EST evidence An indication of whether or not EST evidence is available for the annotation.
- Needs GO An indication of whether or not a new GO term would be needed to characterize the product of the gene model.
- Evidence Assignment of a gene model from a specific track as evidence for the annotation. Choose the name of the gene model and the track from the drop-down menus.
|