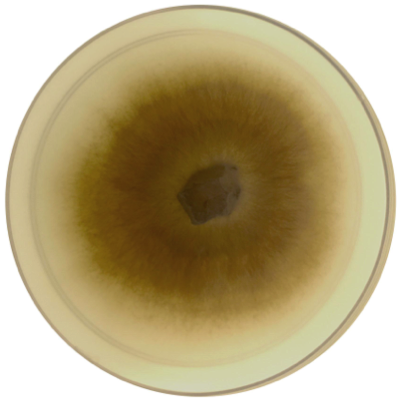
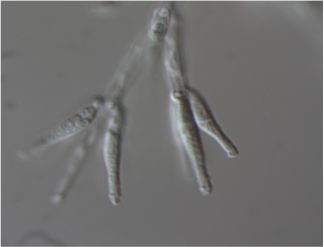
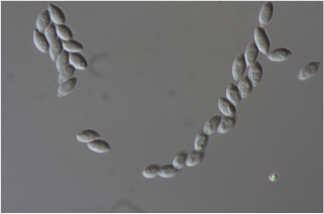
Mariannaea sp. PMI_226 (1113384)
Mariannaea is a monophyletic genus of Ascomycota fungi belonging to phylogenetic group IV within the family Nectriaceae (Samson, 2014; Lombard et al 2015). Like other fungi in the Nectriaceae, Mariannaea spp. are rhizospheric and endophytic associates of any plants in both agricultural and forest soils, and are commonly detected using both culture-based and culture-independent methods (Gottell et al., 2011; Bonito et al., 2014, 2016). Strain PMI_226 was isolated from roots of Quercus alba (Bonito et al., 2016). Based on ITS sequence, PMI_226 is most closely related to other strains (NRRL 62999, CBS 105.66) which were identified as Mariannaea elegans, however additional taxonomic study is still needed to confirm this (Lombard et al., 2015).
This work is supported by the Genomic Science Program (U.S. Department of Energy) Plant Microbe Interfaces (PMI) Scientific Focus Area (http://pmi.ornl.gov) and by the Joint Genome Institute (U.S. Department of Energy) through their Community Sequencing Program (CSP 1974, 1KFG: Deep Sequencing of Ecologically-relevant Dikarya, F. Martin PI).
Researchers
who wish to publish analyses using data from unpublished 1KFG
genomes are respectfully required to contact the PIs and JGI to
avoid potential conflicts on data use and coordinate other
publications with the 1KFG master paper(s).
References
-
Bonito G, Hameed K, Ventura R, Krishnan J, Schadt CW, Vilgalys R. 2016. Isolating a functionally relevant guild of fungi from the root microbiome of Populus. Fungal Ecology 22: 35–42.
-
Bonito G, Reynolds H, Robeson MS, Nelson J, Hodkinson BP, Tuskan G, Schadt CW, Vilgalys R. 2014. Plant host and soil origin influence fungal and bacterial assemblages in the roots of woody plants. Molecular Ecology 23: 3356–3370.
-
Gottel, N.R., Castro, H.F., Kerley, M., Yang, Z.M., Pelletier, D.A., Podar, M., Karpinets, T., Uberbacher, E., Tuskan, G.A., Vilgalys, R., Doktycz, M.J., Schadt, C.W., 2011. Distinct microbial communities within the endosphere and rhizosphere of Populus deltoides roots across contrasting soil types. Appl. Environ. Microbiol. 77 (17), 5934e5944.
-
Lombard,L., van der Merwe,N.A., Groenewald,J.Z. and Crous,P.W. Generic concepts in Nectriaceae. Stud. Mycol. 80, 189-245 (2015)
-
Samson, R.A. 1974. Paecilomyces and some allied Hyphomycetes. Studies in Mycology. 6:1-119
