|
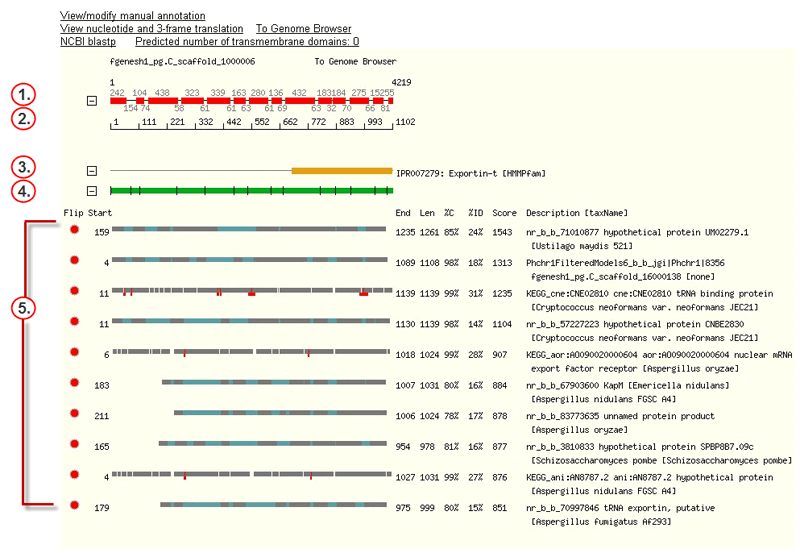
The hit tracks on the Protein page include the following features:
 The JGI-predicted gene (plus strand), showing lengths in base
pairs of predicted exons (red rectangles) and
introns (black lines). Blue rectangles mark
untranslated regions, or UTRs (noncoding exons). The numbers immediately
above the track are exon lengths; those below, intron lengths.
The first and last bases are numbered in the upper left and upper
right. To view the nucleotide sequence, click the gene to open the Gene
Sequence Viewer. The JGI-predicted gene (plus strand), showing lengths in base
pairs of predicted exons (red rectangles) and
introns (black lines). Blue rectangles mark
untranslated regions, or UTRs (noncoding exons). The numbers immediately
above the track are exon lengths; those below, intron lengths.
The first and last bases are numbered in the upper left and upper
right. To view the nucleotide sequence, click the gene to open the Gene
Sequence Viewer.
 A scale showing the numbering of amino acid residues across the
length of the protein sequence derived from the gene. A scale showing the numbering of amino acid residues across the
length of the protein sequence derived from the gene.
 Domain structures in the InterPro database that have alignments
(orange rectangles) with the translated gene.
To collapse these into one additive track (showing alignments
with any of the previously displayed InterPro entities), click the - to the left of the first InterPro
track. The text
for each track links to the European Bioinformatics Institute's
InterPro web page for records with that InterPro ID. The text displays
the InterPro ID number, a description of the InterPro entity, and
in brackets, the algorithm used to determine the match. Domain structures in the InterPro database that have alignments
(orange rectangles) with the translated gene.
To collapse these into one additive track (showing alignments
with any of the previously displayed InterPro entities), click the - to the left of the first InterPro
track. The text
for each track links to the European Bioinformatics Institute's
InterPro web page for records with that InterPro ID. The text displays
the InterPro ID number, a description of the InterPro entity, and
in brackets, the algorithm used to determine the match.
 The protein sequence derived from the predicted gene, with black
lines marking splice junctions. To view the entire amino acid sequence in FASTA
format, click anywhere on the green track. The protein sequence derived from the predicted gene, with black
lines marking splice junctions. To view the entire amino acid sequence in FASTA
format, click anywhere on the green track.
 Alignments (grey rectangles) of proteins from
external databases with the translated gene. The Protein
Alignment Tracks are active areas that can be used to retrieve
alignment details, switch to a flipped view, or link to external
web sites. Alignments (grey rectangles) of proteins from
external databases with the translated gene. The Protein
Alignment Tracks are active areas that can be used to retrieve
alignment details, switch to a flipped view, or link to external
web sites.
|