Getting Started - Tools Overview - KEGG - GO - KOG - Track Editor - Browser - BLAST - Protein Page - Annotation Page
The KOG Browser
EuKaryotic Orthologous Groups (KOG) is a eukaryote-specific version of the Clusters of Orthologous Groups (COG) tool for identifying ortholog and paralog proteins. Where a KOG tool is provided for a JGI-sequenced organism, it provides a way to find JGI-predicted genes by KOG classification or ID.
KOG provides four functional groups, each of which is divided into KOG classifications identified by letters of the alphabet. Within each classification, groups of orthologous or paralogous proteins ("KOGs") are assigned a KOG ID.
With the KOG tool, you can:
- browse through classifications and KOGs.
- search for KOG IDs or descriptions.
- view a list of JGI-predicted genes related to a KOG or classification.
Browse through Classifications and KOGs
The entry page to the KOG tool shows the full list of KOG classifications, ordered by KOG group and letter. Each classification can be expanded to reveal all the KOGs associated with that function. Only one classification can be expanded at a time.
- To expand a KOG classification, click the + symbol next to the classification's letter.
- To collapse a KOG classification, click the - symbol next to the classification's letter or click the + symbol next to a different classification.
For each classification, the number of JGI-predicted genes (gene models) related to that classification in the genome of interest is shown. When you expand a classification, you see the number of gene models related to each KOG. For each KOG group, a total gene count is given as well. The gene counts for classifications and KOGs are links.
- To see a list of gene models that match a classification, click the corresponding number of gene models.
- To see a list of gene models that match a KOG ID, expand the classification and click the number of gene models corresponding to the KOG ID.
Search for KOG IDs or Descriptions
From the KOG entry page, you can begin an advanced search to find genes, transcripts, or proteins by KOG ID or description.
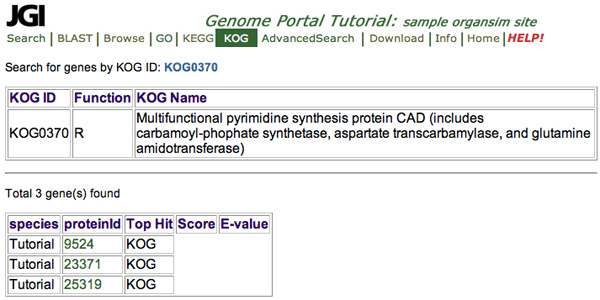
To view the results shown in this picture, from the Search by KOG ID:
- select "Id"
- enter the "KOG0370"
- Click "Submit"
- To search by ID number, click Search by KOG ID at the top of the classification list. You must use the full KOG ID number, including "KOG". To get the correct results when searching for a KOG ID enter the full KOG ID including KOG and any padding zeros, for example enter KOG0370, not "370" or "0370".
- To search by description, click Search by KOG Keyword at the top of the classification list.
- To return to the KOG Browser from the Advanced Search, click the P in the Links column to open the corresponding Protein page, then click the relevant KOG ID or KOG classification name.
View Lists of Predicted Genes
You can view a list of predicted genes (gene models) in a KOG or KOG classification by clicking the corresponding number of gene models on the KOG entry page.
The name of the KOG group appears as the page heading for the list of gene models. Below it is the name of the KOG classification.
- To return to the classification list with the current classification expanded, click the KOG classification heading.
The list of gene models includes: |
|
Prot name |
JGI's name for the predicted gene. |
Prot Id |
The internal JGI identification number for the predicted gene, a link to the Protein page for that gene. |
KOG Id |
The KOG identification number for the KOG to which the gene is annotated. |
KOG description |
The description of the KOG to which the gene is annotated. |
Curated? |
Lists whether or not the model has been manually curated |
To see the Protein page for a specific gene model, click Prot Id.
To see the Annotation page for a specific gene model, click Curated?
At the bottom of the list are buttons for retrieving FASTA sequences or ClustalW alignments.
To retrieve sequence in FASTA format,
- Click the check box next to each gene for which you want to retrieve sequence.
- Click Retrieve FASTA.
To retrieve a ClustalW alignment,
- Click the check box next to each gene that you want to include in the alignment. (You must choose more than one gene.)
- Click Clustalw.
To clear your selections, click Uncheck all.