
Tools Overview
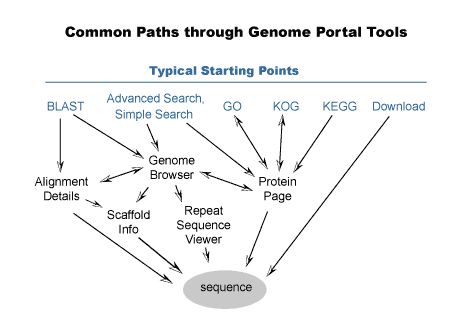
The JGI Genome Portal offers an interface to several bioinformatics tools for studying genomes. Major navigational relationships between the tools are shown below. For each organism sequenced by JGI, a specific subset of tools is available. Throughout a subsite for a given organism, the navigation bar provides links to the tools available for that organism.
Search
This tool allows you to search for InterPro domain predictions or Smith-Waterman alignments to one or more JGI-predicted genes (gene models). You can also jump to a specific JGI gene model.
BLAST
This tool provides an interface to NCBI's Basic Local Alignment Search Tool for queries against JGI genomes. The blastn, tblastx, tblastn, and megablastn options are available. (blastp queries against databases worldwide can be run from the Genome Browser's Protein page.)
Browse
JGI's Genome Browser lets you browse through JGI-predicted genes, view sequences, and study detailed alignments with nucleotide and amino acid sequences from relevant sequence databases.
GO
The Gene Ontology tool uses the Gene Ontology Consortium's controlled vocabulary for organisms to present information about JGI-predicted genes that have automatically assigned GO terms.
KEGG
The KEGG tool uses the Kyoto Encyclopedia of Genes and Genomes and its hierarchies of metabolic and regulatory pathways to present information about JGI-predicted genes that have automatically assigned KEGG terms.
KOG
The KOG tool uses euKaryotic Orthologous Groups, a classification system based on orthologous relationships between genes in eukaryotes, to present information about JGI-predicted genes that have automatically assigned KOG identification numbers.
Advanced Search or Annotation
This search tool provides a more powerful search with features such as wildcards, searches against homologous proteins, and iterative searches. The tool provides links to Genome Browser and Protein page views of results and to locus and trancscript annotation areas.
Download
The Download tool allows you to download a complete JGI genome, predicted proteins, or predicted transcripts for an organism, typically in FASTA format.
Info
The Info page for an organism gives the status of the current version of the genome assembly, tells how the release was created, and details future plans. It also provides funding information and related web links.